Abstract
Key message
A Citrullus amarus mapping population segregating for resistance to Fusarium oxysporum f. sp. niveum race 2 and Papaya ringspot virus was used to identify novel QTL, important for the improvement in watermelon disease resistance.
Abstract
Multiple disease screens of the USDA Citrullus spp. germplasm collection have highlighted the value of Citrullus amarus (citron melon or wild watermelon) as a resource for enhancing modern watermelon cultivars (Citrullus lanatus) with resistance to a broad range of fungal, bacterial and viral diseases of watermelon. We have generated a genetic population of C. amarus segregating for resistance to two important watermelon diseases: Fusarium wilt (caused by the fungus Fusarium oxysporum f. sp. niveum; Fon race 2) and Papaya ringspot virus-watermelon strain (PRSV-W). QTL mapping of Fon race 2 resistance identified seven significant QTLs, with the major QTL representing a novel genetic source of resistance and an opportunity for gene pyramiding. A single QTL was associated with resistance to PRSV-W, which adhered to expectations of a prior study indicating a single-gene recessive inheritance in watermelon. The resistance loci identified here provide valuable genetic resources for introgression into cultivated watermelon for the improvement in disease resistance.




Similar content being viewed by others
References
Ali A, Abdalla O, Bruton B, Fish W, Sikora E, Zhang S, Taylor M (2012) Occurrence of viruses infecting watermelon, other cucurbits, and weeds in the parts of southern United States. Plant Health Progress 13:9. https://doi.org/10.1094/php-2012-0824-01-rs
Branham SE, Levi A, Farnham MW, Patrick Wechter W (2017) A GBS-SNP-based linkage map and quantitative trait loci (QTL) associated with resistance to Fusarium oxysporum f. sp. niveum race 2 identified in Citrullus lanatus var. citroides. Theor Appl Genet 130:319–330. https://doi.org/10.1007/s00122-016-2813-0
Branham SE, Wechter WP, Lambel S et al (2018) QTL-seq and marker development for resistance to Fusarium oxysporum f. sp. niveum race 1 in cultivated watermelon. Mol Breed 38:1–9. https://doi.org/10.1007/s11032-018-0896-9
Branham S, Levi A, Wechter P (2019) QTL mapping identifies novel source of resistance to Fusarium wilt race 1 in Citrullus amarus. Plant Dis. https://doi.org/10.1094/pdis-09-18-1677-re
Broman KW, Sen S (2009) A guide to QTL mapping with R/qtl, vol 46. Springer, New York
Broman KW, Speed T (2002) A model selection approach for the identification of quantitative trait loci in experimental crosses (with discussion). J R Stat Soc B 64(641–656):731–775
Broman KW, Wu H, Sen S, Churchill GA (2003) R/qtl: QTL mapping in experimental crosses. Bioinformatics 19:889–890. https://doi.org/10.1093/bioinformatics/btg112
Brotman Y, Normantovich M, Goldenberg Z et al (2013) Dual resistance of melon to fusarium oxysporum races 0 and 2 and to papaya ring-spot virus is controlled by a pair of head-to-head-oriented nb-lrr genes of unusual architecture. Mol Plant 6:235–238. https://doi.org/10.1093/mp/sss121
Catanzariti A-M, Lim GTT, Jones DA (2015) The tomato I-3 gene: a novel gene for resistance to Fusarium wilt disease. New Phytol 207:106–118. https://doi.org/10.1111/nph.13348
Chandrasekaran J, Brumin M, Wolf D et al (2016) Development of broad virus resistance in non-transgenic cucumber using CRISPR/Cas9 technology. Mol Plant Pathol 17:1140–1153. https://doi.org/10.1111/mpp.12375
Chomicki G, Renner SS (2015) Watermelon origin solved with molecular phylogenetics including Linnaean material: another example of museomics. New Phytol 205:526–532. https://doi.org/10.1111/nph.13163
Cole SJ, Diener AC (2013) Diversity in receptor-like kinase genes is a major determinant of quantitative resistance to Fusarium oxysporum f. sp matthioli. New Phytol 200:172–184. https://doi.org/10.1111/nph.12368
Core Team R (2018) R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna
Gilmour AR, Gogel BJ, Cullis BR, et al. (2009) ASReml user guide release 3.0. VSN International Ltd, Hemel Hemp-stead
Danecek P, Auton A, Abecasis G et al (2011) The variant call format and VCFtools. Bioinformatics 27:2156–2158. https://doi.org/10.1093/bioinformatics/btr330
Diener AC, Ausubel FM (2005) Resistance to Fusarium oxysporum 1, a dominant Arabidopsis disease-resistance gene, is not race specific. Genetics 171:305–321. https://doi.org/10.1534/genetics.105.042218
Elshire RJ, Glaubitz JC, Sun Q et al (2011) A robust, simple genotyping-by-sequencing (GBS) approach for high diversity species. PLoS ONE 6:e19379. https://doi.org/10.1371/journal.pone.0019379
Glaubitz JC, Casstevens TM, Lu F et al (2014) TASSEL-GBS: a high capacity genotyping by sequencing analysis pipeline. PLoS ONE 9:e90346. https://doi.org/10.1371/journal.pone.0090346
Gonsalves D, Tripathi S, Carr JB, Suzuki JY (2010) Papaya ringspot virus. Plant Health Instructor. https://doi.org/10.1094/PHI-I-2010-1004-01
Guner N, Pesic-VanEsbroeck Z, Rivera-Burgos LA, Wehner TC (2018) Inheritance of resistance to Papaya ringspot virus-watermelon strain in watermelon. HortScience 53:624–627. https://doi.org/10.21273/hortsci12944-18
Haley CS, Knott SA (1992) A simple regression method for mapping quantitative trait loci in line crosses using flanking markers. Heredity (Edinb) 69:315–324. https://doi.org/10.1038/hdy.1992.131
Joobeur T, King JJ, Nolin SJ et al (2004) The fusarium wilt resistance locus Fom-2 of melon contains a single resistance gene with complex features. Plant J 39:283–297. https://doi.org/10.1111/j.1365-313X.2004.02134.x
Keinath AP, DuBose V (2009) First report of Fusarium oxysporum f. sp niveum race 2 in South Carolina watermelon fields. Phytopathology 99(6):S63
Kosambi DD (1943) The estimation of map distances from recombination values. Ann Eugen 12:172–175. https://doi.org/10.1111/j.1469-1809.1943.tb02321.x
Lambel S, Lanini B, Vivoda E et al (2014) A major QTL associated with Fusarium oxysporum race 1 resistance identified in genetic populations derived from closely related watermelon lines using selective genotyping and genotyping-by-sequencing for SNP discovery. Theor Appl Genet 127:2105–2115. https://doi.org/10.1007/s00122-014-2363-2
Lander ES, Green P (1987) Construction of multilocus genetic linkage maps in humans. Proc Natl Acad Sci USA 84:2363–2367. https://doi.org/10.1073/pnas.84.8.2363
Lellis AD, Kasschau KD, Whitham SA, Carrington JC (2002) Loss-of-susceptibility mutants of Arabidopsis thaliana reveal an essential role for eIF(iso)4E during potyvirus infection. Curr Biol 12:1046–1051. https://doi.org/10.1016/S0960-9822(02)00898-9
Levi A, Thomas CE, Wehner TC, Zhang X (2001) Low genetic diversity indicates the need to broaden the genetic base of cultivated watermelon. HortScience 36:1096–1101
Levi A, Thies JA, Wechter WP et al (2013) High frequency oligonucleotides: targeting active gene (HFO-TAG) markers revealed wide genetic diversity among Citrullus spp. accessions useful for enhancing disease or pest resistance in watermelon cultivars. Genet Resour Crop Evol 60:427–440. https://doi.org/10.1007/s10722-012-9845-3
Li H, Durbin R (2009) Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25:1754–1760. https://doi.org/10.1093/bioinformatics/btp324
Ling KS, Harris KR, Meyer JD, Levi A, Guner N, Wehner TC, Bendahmane A, Havey MJ (2009) Non-synonymous single nucleotide polymorphisms in the watermelon eIF4E gene are closely associated with resistance to Zucchini yellow mosaic virus. Theor Appl Genet 120(1):191–200
Manichaikul A, Moon JY, Sen Ś et al (2009) A model selection approach for the identification of quantitative trait loci in experimental crosses, allowing epistasis. Genetics 181:1077–1086. https://doi.org/10.1534/genetics.108.094565
Martyn RD (1987) Fusarium oxysporum f. sp. niveum race 2: a highly aggressive race new to the United States. Plant Dis 71:233–236
Meru G, McGregor C (2016a) Genotyping by sequencing for SNP discovery and genetic mapping of resistance to race 1 of Fusarium oxysporum in watermelon. Sci Hortic (Amsterdam) 209:31–40. https://doi.org/10.1016/j.scienta.2016.06.005
Meru G, McGregor CE (2016b) A genetic locus associated with resistance to Fusarium oxysporum f. sp. niveum Race 2 in Citrullus lanatus-type Watermelon. J Am Soc Hortic Sci 141:617–622. https://doi.org/10.21273/JASHS03890-16
Nicaise V, German-Retana S, Sanjuan R et al (2003) The Eukaryotic translation initiation factor 4E controls lettuce susceptibility to the potyvirus lettuce mosaic virus. Plant Physiol 132:1272–1282. https://doi.org/10.1104/pp.102.017855
Ren Y, Jiao Di, Gong G et al (2015) Genetic analysis and chromosome mapping of resistance to Fusarium oxysporum f. sp. niveum (FON) race 1 and race 2 in watermelon (Citrullus lanatus L.). Mol Breed 35:1–9. https://doi.org/10.1007/s11032-015-0375-5
Ruffel S, Dussault MH, Palloix A et al (2002) A natural recessive resistance gene against potato virus Y in pepper corresponds to the eukaryotic initiation factor 4E (elF4E). Plant J 32:1067–1075. https://doi.org/10.1046/j.1365-313X.2002.01499.x
Shimizu M, Fujimoto R, Ying H et al (2014) Identification of candidate genes for fusarium yellows resistance in Chinese cabbage by differential expression analysis. Plant Mol Biol 85:247–257. https://doi.org/10.1007/s11103-014-0182-0
Simons G, Groenendijk J, Wijbrandi J et al (1998) Dissection of the fusarium I2 gene cluster in tomato reveals six homologs and one active gene copy. Plant Cell 10:1055–1068. https://doi.org/10.1105/tpc.10.6.1055
Strange EB, Guner N, Pesic-Vanesbroeck Z, Wehner TC (2002) Plant genetic resources screening the watermelon germplasm collection for resistance to Papaya ringspot virus type-W. Publ Crop Sci 42:1324–1330
Swarts K, Li H, Romero Navarro JA et al (2014) Novel methods to optimize genotypic imputation for low-coverage, next-generation sequence data in crop plants. Plant Genome. https://doi.org/10.3835/plantgenome2014.05.0023
Wechter WP, Kousik C, McMillan M, Levi A (2012) Identification of resistance to Fusarium oxysporum f. sp. niveum race 2 in citrullus lanatus var. citroides plant introductions. HortScience 47:334–338. https://doi.org/10.1002/ird.1717
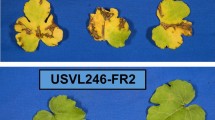
Wechter WP, McMillan MM, Farnham MW, Levi A (2016) Watermelon germplasm lines USVL246-FR2 and USVL252-FR2 tolerant to Fusarium oxysporum f. sp. niveum race 2. HortScience 51:1065–1067
Wehner TC (2008) Watermelon. In Vegetables I:381–418. Springer, New York
Zeng ZB, Kao CH, Basten CJ (1999) Estimating the genetic architecture of quantitative traits. Genet Res 74:279–289. https://doi.org/10.1017/S0016672399004255
Zhang M, Xu JH, Liu G, Yao XF, Li PF, Yang XP (2015) Characterization of the watermelon seedling infection process by Fusarium oxysporum f. sp. niveum. Plant Pathol 64(5):1076–1084
Zheng Y, Wu S, Bai Y et al (2019) Cucurbit genomics database (CuGenDB): a central portal for comparative and functional genomics of cucurbit crops. Nucleic Acids Res 47:D1128–D1136. https://doi.org/10.1093/nar/gky944
Zhou XG, Everts KL, Bruton BD (2010) Race 3, a new and highly virulent race of Fusarium oxysporum f. sp. niveum causing fusarium wilt in watermelon. Plant Dis 94:92–98. https://doi.org/10.1094/PDIS-94-1-0092
Acknowledgements
This research is a part of the Cucurbit Coordinated Agricultural Project (CucCAP) supported by the USDA, National Institute of Food and Agriculture (NIFA), Specialty Crop Research Initiative (SCRI) Grant # 2015-51181-24285, and used resources provided by the SCINet project of the USDA Agricultural Research Service, ARS project number 0500-00093-001-00-D.
Funding
This study was funded, in part, by the US Department of Agriculture (USDA) project number 6080-22000-028-00 and the National Institute of Food and Agriculture, Specialty Crops Research Initiative project number 6080-21000-019-08.
Author information
Authors and Affiliations
Contributions
AL, WPW, SB and KL designed and implemented the experiments. NG, MB and EK developed F2 and F3 population and maintained and supplied PRSV inoculum, WPW, SB, KL and AL phenotyped the population. LM and BC assisted in increasing genetic populations, inoculating and maintaining plants in greenhouse, and in conducting DNA isolation, genotyping and ELISA tests. SB analyzed the data. SB, WPW, KL and AL wrote the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical standards
The experiment conducted complies with the laws of the USA.
Additional information
Communicated by Albrecht E. Melchinger.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Branham, S.E., Patrick Wechter, W., Ling, KS. et al. QTL mapping of resistance to Fusarium oxysporum f. sp. niveum race 2 and Papaya ringspot virus in Citrullus amarus. Theor Appl Genet 133, 677–687 (2020). https://doi.org/10.1007/s00122-019-03500-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-019-03500-3